EAEC Under Stress
Loss of Biofilm Formation
Enteroaggregative Escherichia coli (EAEC) is an emerging foodborne pathogen as a significant diarrheal pathogen in multiple population groups. However, the associated phenotypic changes by which EAEC bacteria survive acid conditions, as it passes through the human digestive track, is not completely understood.
Enteroaggregative E. coli (EAEC) is an important diarrheagenic E. coli, which is increasingly recognized as an emerging pathotype responsible for acute and persistent diarrhea in both developing and developed countries [1-4]. An increasing number of studies have implicated EAEC in endemic diarrhea of infants in both industrialized and developing countries [1], in persistent diarrhea among human immunodeficiency virus/acquired immunodeficiency syndrome patients (HIV) [5]and in traveler’s diarrhea [6]. It has been demonstrated that EAEC can induce growth impairment and malnutrition among children even without diarrhea. The long-term effects of this pathogen in developing countries may be more threatening than the short-term self-limiting diarrhea.
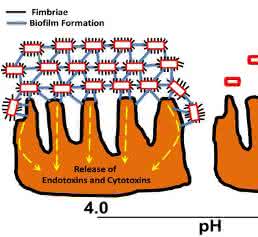
A three-stage model has been proposed for its pathogenesis: [1] characteristic stacked brick-like aggregative adherence (AA) to the intestinal mucosa, HEp-2 cells mediated by 60 MDa plasmid (pAA), also encoding aggregative adherence fimbriae (AAF) for AA phenotype [7-9], hemagglutination (HA) of human erythrocytes, clump and biofilm formation [10-12], [2] increased production and deposition of mucus biofilm, which leads to mucoid stools, malnutrition and persistent colonization [13] and [3] induction of mucosal inflammation with cytokine release, mucosal toxicity and intestinal fluid secretion by enterotoxins and cause destruction of enterocytes [14-18].
Foodborne enteric bacteria including E. coli, E. faecalis, S. typhimurium and H. pylori prefer to live and grow at neutral pH [19]. The pathogenic EAEC has the important property of association with oral-fecal routes of transmission similar to other food-borne pathogens. However, before colonizing the epithelial cells of the intestine, these bacteria pass through the acidic environment of the stomach where the luminal pH is in the range of 1.5-3.5 [20]. Once ingested, these pathogens endure a variety of exposures (acids) in the intestine by colonic microflora. The physiologically triggered pH homeostasis mechanisms include the use of H+ antiport system to maintain internal pH (pHi) at a relatively constant level (pH ~7.6) over a wide range of external pH (pHo) conditions varying from pH 4.3-9.2. Acid dependency of gastric juice is the primary bactericidal barrier against enteric pathogens. The mechanism employed by EAEC to sense and respond to acidic pH has not been elucidated so far. Therefore, an attempt has been made in EAEC to study the effect of variable pH conditions (acid stress), simulating natural route of infection (pH 1.5-5.5). This work will enable us to identify the associated phenotypic changes responsible for its adaptation and survival under acid stress conditions in order to correlate with EAEC-induced pathogenesis.
Materials and Methods
Bacterial strain and growth conditions
EAEC (T8) strain was procured from National Institute of Cholera and Enteric Diseases (NICED), Kolkata. It was characterized both biochemically and phenotypically (clump formation, hemagglutination assay and HEp-2 adherence assay) for the study. EAEC (T8) was grown on Luria Agar (LA; Sigma-Aldrich, USA) plates for 16h at 37°C.
Acid stress (Low pH): In order to study growth under acid stress (inorganic) conditions [21], the pH of minimal media (M9) was adjusted with 1 N HCl to variable pH (3.0-5.5). EAEC (T8) was grown in shaker (Innova, 4230 Refrigerated Incubator shaker, New Brunswick Scientific Edison, NJ USA) at 37°C with shaking (215 rpm).
Growth curve analysis: For growth curve analysis, an inoculation or starter culture is used. Bacterial cultures should always be grown from a single colony picked from a freshly streaked selective plate. Subculturing directly from glycerol stocks, agar stabs and liquid cultures and inoculation from plates that have been stored for a long time is a poor microbiological practice. Briefly, a single colony of EAEC (T8) was inoculated in 10 ml of LB broth (pH 7.4) and was allowed to grow overnight at 37°C with shaking at 215 rpm. Using a flask with a volume of at least four times greater than the volume of medium, the starter culture (from 10 ml) was diluted 1:500 into the pre-warmed media of M9 (variable pH) (400ml) and incubated under similar conditions (37°C; 215 rpm). Every hour, 1 ml samples were recorded and absorbance (OD 600 nm) was measured in a spectrophotometer (Kontron 860 Spectrophotometer, Netherlands) till OD 600 nm reaches approximately 2.0.
Cell line and cell culture: The HEp-2 cell line (source: human pharynx) was obtained from national centre for cell sciences (NCCS), Pune, India. The HEp-2 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (pH 7.4) and maintained at 37°C with humidified 5% CO2 in 25 cm2 tissue culture flask (Greiner, USA) with antibiotics Penicillin G (100 U/ml) and Streptomycin (100 µg/ml). As a routine cell passage, 10% fetal calf serum (FCS; Sigma-Aldrich) was used.
HEp-2 adherence assay
HEp-2 adherence assay for EAEC (T8) characterization was done by the method described by [22] with some modifications. The effect of media pH on the adherence of EAEC (T8) infected HEp-2 cells (in vitro) was also studied. Cells were grown to 50% to 70% confluence as monolayers in a 6-well flat-bottom tissue culture plate. The HEp-2 cells were washed three times with phosphate buffer saline (PBS) and 2 ml of fresh DMEM media set with 1 N HCl with desired pH (filter sterilized) was added along with 2% FCS and 0.5% D-mannose without antibiotics to the 6-well plate. To this, EAEC (T8) grown overnight at 37°C (215 rpm) was inoculated (25 µl) in the plate and incubated at 37°C overnight with 5% CO2. Following incubation, the cells were washed three times with PBS, fixed with 100% methanol and stained with 2.5% Giemsa for 15 min. The adherence patterns were examined under 40-X magnification and finally photographed at 100-X magnification with digital camera (Carl Zeiss, Sony, Japan) with oil immersion in a light microscope (Leica MPS32, USA).
Propidium iodide (PI) exclusion assay
The adherent cell survival at various pH was also determined (in vitro) on HEp-2 cells. For this, FACS assay was carried out with slight modifications[23]. After incubation of HEp-2 cells with EAEC (T8), cells were trypsinized and removed with PBS and EDTA (0.02%). The cells were harvested by centrifugation (4000 rpm for 20 min) and washed twice with PBS (pH 7.4). Finally, the pellet was resuspended in PBS containing PI and RNase A in dark for 2h at 4°C. For the detection of cell survival, 1-X FACS sheath buffer was added for sample analysis (Becton Dickinson, CA, USA, FACS Calibur equipped with 15 MW, 488 nm air cooled argon laser) using Cell Quest software. DNA of HEp-2 cells cultured in DMEM media (pH 4.0 and 7.4) for 3h incubation was extracted to examine any DNA fragmentation. The cells with degraded DNA incorporate less PI than the cells with intact DNA.
Clump formation test: The clump formation test, which is specific for EAEC (T8) was performed as described by [24]. Briefly, EAEC (T8) was subcultured on LA and then further inoculated (1:500) in 5 ml of each of LB in duplicate tubes. One set of tubes was incubated in the stationary position and the other set in an orbital shaker incubator at 37°C. The formation of a clump or a pellicle as a ring at the side of the test tube at the end of incubation (20h) was regarded as a positive result.
Hemagglutination assay
Hemagglutination (HA) was performed with human type A erythrocytes as described by [25] with minor modifications. EAEC (T8) grown overnight in tryptone soya broth under static conditions was pelleted, washed and resuspended in PBS. Twenty five microliters of the suspension was mixed briefly with an equal volume of 3% (v/v) washed erythrocyte suspension in PBS containing 1% D-mannose and was allowed to stand at 4°C to 8°C for 30 min. HA was scored either as positive or negative for umbrella and button formation respectively.
Biofilm assay
(i) Quantitative method: To assess biofilm formation quantitatively [26], an overnight culture of EAEC (T8) in LB was subcultured (1:500) in prewarmed DMEM containing 0.45% glucose in 96-well flat-bottom microtiter polystyrene plates (Costar 3595; Corning Inc., Corning, NY). Plates were incubated at 37°C for 24h and the culture medium was then decanted and the plates were washed twice with 200 µl of sterile double distilled water (ddw) to remove the loosely and unbound cells. The adherent bacteria were stained with 50 µl of 0.1% crystal violet for 15 min and then rinsed twice with ddw (200 µl). The bound dye was extracted from the stained cells by washing with 99% ethanol (200 µl). The biofilm was quantified in duplicate for each sample and absorbance (OD 570 nm) of the solution was measured in an enzyme linked immunosorbent assay reader (ELISA reader, Bio-Rad).
(ii) Microscopic method: Microtiter plate assays were performed as described by [27]. To assess biofilm formation, an overnight culture of EAEC (T8) in LB was subcultured (1:500) in prewarmed high glucose of DMEM containing 0.45% glucose in 6-well flat-bottom microtiter polystyrene plates. Plates were incubated at 37°C for 24h and culture medium was then decanted and the plates were washed twice with 200 µl of sterile ddw to remove the loosely and unbound cells. The adherent bacteria were stained with 50 µl of 0.1% crystal violet for 15 min and were rinsed twice with 200 µl of water. The biofilm formed on the plate was finally photographed at 100-X magnification with digital camera (Carl Zeiss, Sony, Japan) with oil immersion in a light microscope (Leica MPS32, USA).
Transmission electron microscopy
For electron microscopy, modified procedure of [28] was followed. EAEC (T8)-infected HEp-2 cells after 3h of incubation at pH 4.0 and control (7.4) were trypsinized and centrifuged at (4000 rpm for 20 min) and were washed in 3% buffered glutaraldehyde (made in Sorenson’s buffer, pH 7.2). To the pellet, after low speed centrifugation, 1 ml of 3% glutaraldehyde was added and centrifuged (4000 rpm for 10 min). The supernatant was decanted and the cells were fixed by adding 3% buffered glutaraldehyde for 4h and the pellet was washed with the same buffer. The pellet was post fixed in 1% osmium tetraoxide, embedded in 1% agar. The embedded sample was dehydrated in graded series of alcohol and finally embedded in Araldite resin. It was then sectioned and post stained with uranyl acetate and lead. The samples were viewed with a transmission electron microscope (Zeiss, 906, Germany) to determine the morphological changes, if any associated with low pH.
Results
Acid stress suppresses growth of EAEC (T8)
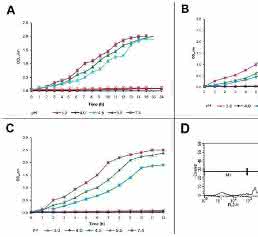
The growth pattern of EAEC (T8) was studied in vitro by adjusting M9 as well as LB media pH values (one or two points) on either side of the mean gastric pH with concentrated HCl, ranging from pH 3.0-7.4. At pH 4.0 in LB media EAEC (T8) demonstrated viable growth [29]. The minimum lowest pH in M9 media where EAEC (T8) was able to grow linearly was pH 4.5. EAEC (T8) demonstrated similar growth pattern at pH 4.5 as well as 5.5, which is the mild acidic pH (mimicking intestinal pH) compared to pH 7.4 (control) in M9 media (Figure 1A). The log phase at pH 4.5 and 5.5 was attained after 7 and 6h respectively. However, the log phase growth at control pH 7.4 was achieved in approximately 4h (Figure 1A).
It is known that decarboxylases play an important role in enterics during acid stress, however, their expression is induced only in the presence of specific amino acids Therefore, the growth pattern of EAEC (T8) was studied at various pH in M9 media with supplements lysine and arginine (0.12%). We demonstrate that the arginine and lysine supplements in M9 media did not significantly alter the growth rate and EAEC (T8) was able to grow up to pH 4.5. However, at pH 3.0 and 4.0 in M9 media with either of the supplements, EAEC (T8) did not grow (Figure 1B and 1C). The log phase in M9 media with lysine and arginine supplements at pH 4.5 reached in 7h and 6h, respectively. EAEC (T8) at pH 5.5 in minimal media reached log phase in 5h with lysine (Figure 1B) and 4h with arginine supplements (Figure 1C). However, at pH 7.4 (control) doubling time was same of EAEC (T8) with the supplements (Figure 1B and 1C). However, the minimum growth pH 4.0 was observed in LB media where EAEC (T8) demonstrates viability was taken for the phenotypic characterization.
Aggregative adherence is a characteristic property of EAEC[30]. Adherence to host HEp-2 cells and percentage survival in vitro was determined at pH 4.0 and 7.4 (control) by FACS analysis. The survival rate of EAEC (T8)-infected with HEp-2 cells at pH 7.4 was approximately 84% (Figure 1D; right panel), whereas at pH 4.0 the survival rate 54% (Figure 1D; left panel). The possibility exists that the reduction in cell survival number could be due to decreased proliferation or increased cell death. Further, DNA was prepared from EAEC (T8)-infected HEp-2 cells at both pH 4.0 and 7.4. At pH 4.0, DNA demonstrated smear characteristics (data not shown).
Loss of clump formation of EAEC (T8)
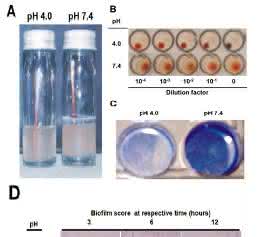
We demonstrate that clump formation appeared as a thin ring varying in breadth from 0.4 cm to 0.8 cm at the top of the liquid culture at pH 7.4 after 20h of incubation in LB and remained intact to the test tube as a ring under both static as well as shaking conditions (Figure 2A; right panel). However, EAEC (T8) grown at pH 4.0 did not produce clumps at 37°C even under static conditions (Figure 2A; left panel).
Loss of hemagglutination of EAEC (T8)
At pH 7.4 in EAEC (T8), the umbrella shaped hemagglutination was observed and even at the highest dilution (10-4), hemagglutination was intact (Figure 2B; bottom)]. In contrast, button-shaped hemagglutination was ob-served in EAEC (T8) at pH 4.0 (Figure 2B; top).
Loss of biofilm formation of EAEC (T8)
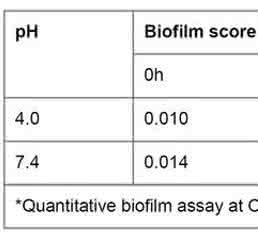
Biofilm formation at pH 4.0 was determined both quantitatively and microscopically and was found to be significantly less than at pH 7.4 (Figure 2C, left panel). The quantitative biofilm score (OD 570 nm) was ≤0.1 at pH 4.0 in EAEC (T8). Therefore, even after 12h of incubation, the score was only 0.112. However, at pH 7.4, the biofilm formation was found to be at least four-fold more than the pH 4.0, with a score of 0.449 (Table 1). Further, on microscopic image analysis, the biofilm score was negligible at pH 4.0 (<+1) even after 12h of incubation (Figure 2D; top) However, in control (pH 7.4), EAEC (T8) demonstrated a linear increase in the rate of biofilm formation, with a score of +1, +2 and +3 respectively after 3, 6 and 12h of growth (Figure 2D, bottom).
Loss of typical aggregative adherence pattern of EAEC (T8)
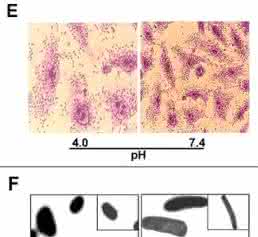
During acid stress or under low pH of 4.0, the phenotypic properties of EAEC (T8) bacteria were altered. Diffusely adherent pattern was observed under acid stress pH 4.0 (Figure 2E; left panel) in contrast to the typical „stacked brick“ aggregative adherence in control pH 7.4 (Figure 2E; right panel).
Loss of characteristic rod-shape of EAEC (T8)
At pH 7.4 (control), EAEC (T8) bacteria exhibited characteristic rod shapes with average dimensions of 4.5 mm × 0.8 mm and even after 3h of incubation with HEp-2 cells, there was no alteration in the cell morphology. However, within 3h of incubation at pH 4.0 in DMEM media, rod-shaped cells appeared as coccoid or spheroid forms with average dimensions of 2.5 mm × 1.4 mm (Figure 2F; left panel). These acid stressed cells became morphologically shorter and reduced approximately to half of the normal sized cells (Figure 2F; right panel).
Discussion
Enteroaggregative Escherichia coli (EAEC), a diarrheagenic E. coli is implicated in persistent pediatric diarrhea in both developing and industrialized countries (REF). The pathogenesis of EAEC infection is not well understood due to the genetic and phenotypic heterogeneity. Similar to any other enteric bacteria, EAEC is also exposed to low pH conditions during their migration from stomach (pH 1.5) to intestine (pH 5.5). Acid resistance (AR) could be an indicator of virulence as only acid resistant strains are able to survive the human stomach passage and causes infection (REF). Therefore, in this study we looked the effect of changing pH on EAEC and its ability to survive, adhere and infect.
In order to monitor the AR in the clinical isolate EAEC (T8), growth pattern was studied in various M9 as well as LB media. The acid tolerance of EAEC (T8) was found to be media-dependent. In the present study, EAEC (T8) demonstrated very similar growth pattern at acidic pH 4.5 in both M9 and M9 with amino acid supplements (Figure 1A-1C). However we have demonstrated that in LB broth, the lowest pH at which EAEC (T8) demonstrated linear growth was pH 4.0 [29]. In contrast, at pH 3.0 even after 3 days in the shaker (215 rpm), there was no growth (data not shown). The growth at pH 4.0 in LB was almost three fold slower than the control pH (7.4) [29]. EAEC (T8) was sensitive to killing in acidified media at pH 3.0 in LB while in minimal (M9) media at pH 4.0. It might be correlated to the earlier findings where LB media reported to be a rich source for culturing of enteric bacteria [31,32] while in other media, they failed to initiate rapid growth. The capacity to invoke acid tolerance response (ATR) in S. mutans on teeth ensures increased survival at pH 4.3 [33]. Similar results demonstrated that in defined medium (Sauton), the growth of M. tuberculosis was completely absent at pH 6.0 [34].
The disparity in results is not known, however the possibility might exist that is due to various media used since the sensitivity to extreme pH ranges can be masked in complex media. However, EAEC encounters very low pH (1.5) in stomach during ingestion, which might be correlated well with the previous reports that some strains of E. coli are able to survive at pH values as low as 2.5, however, it does not grow at pH values less than 4.4 [35,36]. Growth inhibition occurs as a result of both lower pHi and the ability of anions to inhibit metabolism. E. faecalis demonstrated growth within a range of pHo 4.5-9.5[37]. N. gonorrhoeae had survival peak at pH 4.6-4.7 [38] and Y. enterocolitica demonstrate survival peak at pH 4.5[39]. Enterohemorrhagic E. coli (EHEC) can survive pH 2.0 for 5h [40] whereas nontoxigenic strain can survive pH 3.0 only [41]. It has been demonstrated that these bacteria cannot survive under extreme acidic conditions and therefore, food could provide a protective effect to acidsensitive barrier by facilitating their survival under extreme acidic conditions. During infection in stomach, the bacteria is already in stationary phase (non-dividing) and it is well reported that the survival potential of stationary phase or acid-adapted cells is greater than that of exponentially growing cells over the initial period of acid challenge [41]. So, once induced, the AR system will remain active until cells reenter log phase [42]. Therefore, the growth in vivo at pH 2.4 might compensate the survival of EAEC (T8) at pH 4.0 in vitro. Thus, the bacteria might be adapted when it encounters low pH in stomach.
In our study, the phenotypic properties of EAEC including typical aggregative adherence (AAt), hemagglutination (HA) and biofilm formation were not observed at pH 4.0 (Figure 2B-2E). As it is well known that EAEC characterized by “stacked-brick” adherence phenotype, which is mediated by aggregative adherence fimbriae (AAFs) encoded by 60 MDa megaplasmid [30]. The transfer of the megaplasmid from EAEC to a laboratory E. coli strain transferred the property of aggregative adherence [43]. One study reported that a EHEC strain of serotype O103:H2 demonstrated loss of this megaplasmid coincided with reduced adhesion to cultured epithelial cells [44] while the other EHEC strain of serotype O5:H did not demonstrate any effect on adhesion. The invasion plasmid antigens (Ipa proteins) in Shigella, required for invasion of the colonic and rectal epithelial cells [45] were downregulated at acidic pH, which might explain that Shigella affects the lower gut where cellular invasion occurs and the acidic environment of the stomach prevents expression of such virulence genes [46]. CadA, a lysine decarboxylase modulates expression of the intimin, an outer membrane adhesin involved in pathogenesis; negatively regulate virulence in several enteric pathogens and in EHEC strains. An inactivated cadA in EHEC did not possess lysine decarboxylation activity and was hyperadherent to tissue cultured cells by nearly twofold [47]. Disruption of the intimin encoding EAE gene in the cadA mutant significantly reduced its adherence to tissue-cultured cells [47]. Two putative adhesins, flagella and F9 fimbria, were upregulated in the cadA mutant, suggestive of their association with adherence in the absence of the Cad regulatory mechanism [47]. In the present study, the loss of AAt phenotype of EAEC (T8) at pH 4.0 might be also correlated with the downregulation of genes encoded by megaplasmid. Our results demonstrated DNA smearing of EAEC (T8) when infected with HEp-2 cells at pH 4.0 is consistent with the earlier reports where acid stress induced DNA damage [48].
The AAt pattern was observed of EAEC (T8) at control pH (7.4), however, at pH 4.0, it demonstrates diffuse adherence (DA) (Figure 2E). The expression of 34 kDa OMP was high and 41 kDa and 48 kDa OMP were newly synthesized at pH 4.0 in EAEC (T8), which might be involved in DA adherence [29]. A 30 kDa to 43 kDa OMP have role in aggregative adherence and hemagglutination [49]. The fimbriae adhesin (18 kDa) from an Indian strain of EAEC demonstrated MRHA and HEp-2 cell adherence [50]. Moreover, vaccines inducing anti-adhesin immunity to inhibit bacterial adherence of K88 fimbrial ETEC to porcine small intestinal enterocytes [51]. It is well reported that a 100 kDa OMP [52] and fibrillar adhesin [53] are associated with the DA phenotype pattern in EPEC. These findings correlate well with the altered OMP expression and some of the OMP might be downregulated at acidic pH in EAEC (T8), might play role in EAEC (T8) survival [29]. There- fore, acidic exposure in the stomach might create a new phenotype overall, which causes infection only in the intestine after adapting and surviving at low pH of stomach.
Besides, the aggregative adherence, the clump or scum formation was also completely abolished in EAEC (T8) at pH 4.0 (Figure 2A). These results were similar to the earlier findings where 100% correlation between scum formation and the aggregative adherence pattern has been reported in EAEC [10]. The most dominant AAt patterns were mostly associated with the AAF-related gene sequences or their regulator, the aggR gene. It is reported that aggR-positive EAEC strains with the AAt pattern demonstrated significantly stronger biofilm formation (OD570 0.72) than did those with the atypical AA pattern (OD570 0.36) [26]. These results are consistent with our findings that AAt pattern (pH 7.4) demonstrated high score (OD570 0.72) for biofilm formation than the DA pattern at pH 4.0 (OD570 0.36). Thus, EAEC (T8) exhibiting other types of adherence, might be devoid of any of the AAF-related genes.
In the present study, the cell morphology by transmission electron microscopy (TEM) demonstrated that the flagellum was intact in EAEC (T8) cell, however, the cell size was reduced to almost half at low pH (4.0) (Figure 2F). V. parahaemolyticus are normally rod shaped cells but appeared as irregular shapes and finally became coccoid or spheroid forms and reduced half in a week during starvation [54] for 1 week and were even without a flagellum. It has been proposed that cell size re-duction during starvation is a survival strategy for minimizing cell maintenance requirements and enhancing substrate uptake due to a high surface-volume ratio [55].
The overall complexity of the stress response in enteric bacteria is probably greatly underestimated, since intracellular growth produces global patterns substantially various from those predicted by stress experiments under defined conditions [56]. It appears that there are indeed numerous acid survival mechanisms; some have long-term dramatic effect while others have more subtle yet significant consequences. We observed changes in the morphological features might help EAEC (T8) to survive under acidic conditions in the stomach. Our previous studies has demonstrated that stationary phase gene regulator rpoS plays very important role in the regulation of acid shock proteins induced at low pH 4.0 as the rpoS mutant didn’t grow at pH 4.0 [29]. The apparent relevance of growth conditions to the changes in the phenotypic characteristics of EAEC (T8) provided us the information regarding the effect of environmental stresses on the virulence of EAEC. Therefore, the altered phenotypic changes might hold well in adaptation and protection of EAEC (T8) through low pH in stomach. Further studies on the effect of bile salts and pancreatic juices found in small intestine on EAEC (T8) virulence would shed more light on EAEC survival.
Conclusions
EAEC (T8) demonstrated media-dependent growth rate. The re-duction in cell survival of EAEC (T8) grown in vitro with HEp-2 cells at pH 4.0 could be due to decreased proliferation or increased cell death, which could be correlated well with the possibility of infection in intestine (in vivo) and not in stomach (pH 1.5).
In addition, EAEC (T8) also demonstrated changes in its phenotypic characters during acid stress (pH 4.0), which were evident by lack of the clump formation and hemagglutination. The biofilm formation at pH 4.0 was negligible, which is a deviation from the three-stage model proposed for EAEC pathogenesis. The lack of typical aggregative adherence pattern at pH 4.0 of EAEC (T8) might be related with the changes in cell morphology (Figure 3).
Funding
This work was funded in part by institutional support from Morehouse School of Medicine and U54 CA118638 (to P.K.) and institutional support from Scott & White Memorial Hospital and Clinic, Texas A&M Health Science Center College of Medicine, the Central Texas Veterans Health Administration, the US National Institutes of Health grant RO1CA91889, an Endowment from the Cain Foundation and University of Dammam (to A.A.). The authors alone are responsible for the content and writing of the paper. The authors thank Dr. Anuradha Chakraborti for providing critical reagents and Ms. Meenakshi Kaundal for her expert technical assistance. The authors have declared that no competing interests exist.
© 2017 Kaur P, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Citation: Kaur P, Asea A (2017) Loss of Biofilm Formation in an Emerging Foodborne Pathogen Enteroaggregative Escherichia coli (EAEC) Under Acid Stress. JCEST 8: 260. doi:10.4172/2157-7013. 1000260
Punit Kaur
Department of Microbiology, Biochemistry and Immunology
Morehouse School of Medicine
Atlanta, GA, USA
and Department of Experimental RadHiation Oncology
University of Texas MD
Anderson Cancer Center
Houston, Texas, USA
Alexzander Asea
Department of Neuroscience Research and Deanship for Scientific Research
University of Dammam
Dammam, Saudi Arabia
Corresponding author
Dr. Punit Kaur
Email: [email protected], [email protected]
References
[1] Huang DB, Koo H, DuPont HL (2004) Enteroaggregative Escherichia coli: An Emerging Pathogen. Curr Infect Dis Rep 6: 83-86.
[2] Harada T, Hiroi M, Kawamori F, Furusawa A, Ohata K, et al. (2007) A food poisoning diarrhea outbreak caused by enteroaggregative Escherichia coli serogroup O126:H27 in Shizuoka, Japan. Jpn J Infect Dis 60: 154-155.
[3] Okhuysen PC, Dupont HL (2010) Enteroaggregative Escherichia coli (EAEC): A cause of acute and persistent diarrhea of worldwide importance. J Infect Dis 202(4): 503-505.
[4] Rasko DA, Webster DR, Sahl JW, Bashir A, Boisen N (2011) Origins of the E. coli strain causing an outbreak of hemolytic-uremic syndrome in Germany. N Engl J Med 365: 709-717.
[5] Mayer HB, Wanke CA (1995) Enteroaggregative Escherichia coli as a possible cause of diarrhea in an HIV-infected patient. N Engl J Med 332: 273-274.
[6] Adachi JA, Ericsson CD, Jiang ZD, DuPont MW, Pallegar SR (2002) Natural history of enteroaggregative and enterotoxigenic Escherichia coli infection among US travelers to Guadalajara, Mexico. J Infect Dis 185: 1681-1683.
[7] Boisen N, Struve C, Scheutz F, Krogfelt KA, Nataro JP (2008) New adhesin of enteroaggregative Escherichia coli related to the Afa/Dr/AAF family. Infect Immun 76: 3281-3292.
[8] Chaudhuri RR, Sebaihia M, Hobman JL, Webber MA, Leyton DL (2010) Complete genome sequence and comparative metabolic profiling of the prototypical enteroaggregative Escherichia coli strain 042. PLoS One 5: e8801.
[9] Boll EJ, Struve C, Sander A, Demma Z, Nataro JP (2012) The fimbriae of enteroaggregative Escherichia coli induce epithelial inflammation in vitro and in a human intestinal xenograft model. J Infect Dis 206: 714-722.
[10] Bhardwaj R, Majumdar S, Ganguly NK, Taneja N, Dutta S (2006) Characterization of adhesin variants in Indian isolates of enteroaggregative Escherichia coli. FEMS Microbiol Lett 258: 274-283.
[11] Macleod SM, Stickler DJ (2007) Species interactions in mixed-community crystalline biofilms on urinary catheters. J Med Microbiol 56: 1549-1557.
[12] Wang X, Lunsdorf H, Ehren I, Brauner A, Romling U (2010) Characteristics of biofilms from urinary tract catheters and presence of biofilm-related components in Escherichia coli. Curr Microbiol 60: 446-53.
[13] Harrington SM, Sheikh J, Henderson IR, Ruiz-Perez F, Cohen PS (2009) The Pic protease of enteroaggregative Escherichia coli promotes intestinal colonization and growth in the presence of mucin. Infect Immun 77: 2465-2473.
[14] Fagundes-Neto U, De Martini-Costa S, Pedroso MZ, Scaletsky IC (2000) Studies of the small bowel surface by scanning electron microscopy in infants with persistent diarrhea. Braz J Med Biol Res 33: 1437-1442.
[15] Muniesa M, Hammerl JA, Hertwig S, Appel B, Brussow H (2012) Shiga toxin-producing Escherichia coli O104:H4: A new challenge for microbiology. Appl Environ Microbiol 78: 4065-4073.
[16] Nataro JP (2011) Outbreak of hemolytic-uremic syndrome linked to Shiga toxin-producing enteroaggregative Escherichia coli O104:H4. Pediatr Res 70: 221.
[17] Frank C, Werber D, Cramer JP, Askar M, Faber M (2011) Epidemic profile of Shiga-toxin-producing Escherichia coli O104:H4 outbreak in Germany. N Engl J Med 365: 1771-1780.
[18] Strauman MC, Harper JM, Harrington SM, Boll EJ, Nataro JP (2010) Enteroaggregative Escherichia coli disrupts epithelial cell tight junctions. Infect Immun 78: 4958-4964.
[19] Gorden J, Small PL (1993) Acid resistance in enteric bacteria. Infect Immun 61: 364-367.
[20] Verdu E, Viani F, Armstrong D, Fraser R, Siegrist HH (1994) Effect of omeprazole on intragastric bacterial counts, nitrates, nitrites, and Nitroso compounds. Gut 35: 455-460.
[21] Foster JW, Hall HK (1990) Adaptive acidification tolerance response of Salmonella typhimurium. J Bacteriol 172: 771-778.
[22] Cravioto A, Tello A, Navarro A, Ruiz J, Villafan H (1991) Association of Escherichia coli HEp-2 adherence patterns with type and duration of diarrhoea. Lancet 337: 262-264.
[23] Oberdoerster J, Kamer AR, Rabin RA (1998) Differential effect of ethanol on PC12 cell death. J Pharmacol Exp Ther 287: 359-365.
[24]. Albert MJ, Qadri F, Haque A, Bhuiyan NA (1993) Bacterial clump formation at the surface of liquid culture as a rapid test for identification of enteroaggregative Escherichia coli. J Clin Microbiol 31: 1397-1399.
[25] Yamamoto T, Endo S, Yokota T, Echeverria P (1991) Characteristics of adherence of enteroaggregative Escherichia coli to human and animal mucosa. Infect Immun 59: 3722-3739.
[26] Wakimoto N, Nishi J, Sheikh J, Nataro JP, Sarantuya J (2004) Quantitative biofilm assay using a microtiter plate to screen for enteroaggregative Escherichia coli. Am J Trop Med Hyg 71: 687-690.
[27] O‘Toole GA, Kolter R (1998) Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol Microbiol 30: 295-304.
[28] Root RK, Isturiz R, Molavi A, Metcalf JA, Malech HL (1981) Interactions between antibiotics and human neutrophils in the killing of staphylococci. J Clin Invest 67: 247-259.
[29] Kaur P, Chakraborti A (2010) Proteome analysis of a food borne pathogen Enteroaggregative Escherichia coli under acid stress. J Proteomics Bioinform 3: 10-19.
[30] Nataro JP, Yikang D, Yingkang D, AggR WK (1994) A transcriptional activator of aggregative adherence fimbria I expression in enteroaggregative Escherichia coli. J Bacteriol 176: 4691-4699.
[31] Chervaux C, Ehrlich SD, Maguin E (2000) Physiological study of Lactobacillus delbrueckii subsp. bulgaricus strains in a novel chemically defined medium. Appl Environ Microbiol 66: 5306-5311.
[32] Partanen L, Marttinen N, Alatossava T (2001) Fats and fatty acids as growth factors for Lactobacillus delbrueckii. Syst Appl Microbiol 24: 500-506.
[33] Bowden GH, Hamilton IR (1998) Survival of oral bacteria. Crit Rev Oral Biol Med 9: 54-85.
[34] Hackam DJ, Rotstein OD, Zhang W, Gruenheid S, Gros P (1998) Host resistance to intracellular infection: Mutation of natural resistance associated macrophage protein 1 (Nramp1) impairs phagosomal acidification. J Exp Med 188: 351-364.
[35] Small P, Blankenhorn D, Welty D, Zinser E, Slonczewski JL (1994) Acid and base resistance in Escherichia coli and Shigella flexneri: role of rpoS and growth pH. J Bacteriol 176: 1729-1737.
[36] Benjamin MM, Datta AR (1995) Acid tolerance of enterohemorrhagic Escherichia coli. Appl Environ Microbiol 61: 1669-1672.
[37] Kobayashi H (1985) A proton-translocating ATPase regulates pH of the bacterial cytoplasm. J Biol Chem 260: 72-76.
[38] Pettit RK, McAllister SC, Hamer TA (1999) Response of gonococcal clinical isolates to acidic conditions. J Med Microbiol 48: 149-156.
[39] De Koning-Ward TF, Robins-Browne RM (1995) Contribution of urease to acid tolerance in Yersinia enterocolitica. Infect Immun 63: 3790-3795.
[40] Brown JL, Ross T, McMeekin TA, Nichols PD (1997) Acid habituation of Escherichia coli and the potential role of cyclopropane fatty acids in low pH tolerance. Int J Food Microbiol 37: 163-173.
[41] Jordan KN, Oxford L, O‘Byrne CP (1999) Survival of low-pH stress by Escherichia coli O157:H7: correlation between alterations in the cell envelope and increased acid tolerance. Appl Environ Microbiol 65: 3048-3055.
[42] Lin J, Smith MP, Chapin KC, Baik HS, Bennett GN, et al. (1996) Mechanisms of acid resistance in enterohemorrhagic Escherichia coli. Appl Environ Microbiol 62: 3094-4100.
[43] Vial PA, Robins-Browne R, Lior H, Prado V, Kaper JB, et al. (1988) Characterization of enteroadherent-aggregative Escherichia coli, a putative agent of diarrheal disease. J Infect Dis 158: 70-79.
[44] Hall GA, Dorn CR, Chanter N, Scotland SM, Smith HR, et al. (1990) Attaching and effacing lesions in vivo and adhesion to tissue culture cells of Vero-cytotoxin-producing Escherichia coli belonging to serogroups O5 and O103. J Gen Microbiol 136: 779-786.
[45] Yang F, Yang J, Zhang X, Chen L, Jiang Y (2005) Genome dynamics and diversity of Shigella species, the etiologic agents of bacillary dysentery. Nucleic Acids Res 33: 6445-6458.
[46] Dorman CJ, McKenna S, Beloin C (2001) Regulation of virulence gene expression in Shigella flexneri, a facultative intracellular pathogen. Int J Med Microbiol 291: 89-96.
[47] Vazquez-Juarez RC, Kuriakose JA, Rasko DA, Ritchie JM, Kendall MM (2008) CadA negatively regulates Escherichia coli O157:H7 adherence and intestinal colonization. Infect Immun 76: 5072-5081.
[48] Foster JW (1999) When protons attack: microbial strategies of acid adaptation. Curr Opin Microbiol 2: 170-174.
[49] Wai SN, Takade A, Amako K (1996) The hydrophobic surface protein layer of enteroaggregative Escherichia coli strains. FEMS Microbiol Lett 135: 17-22.
[50] Grover V, Ghosh S, Chakraborti A, Majumdar S, Ganguly NK (2007) Galactose-specific fimbriae adhesin of enteroaggregative Escherichia coli: a possible aggregative factor. Curr Microbiol 54: 175-179.
[51] Zhang C, Zhang W (2010) Escherichia coli K88ac fimbriae expressing heat-labile and heat-stable (STa) toxin epitopes elicit antibodies that neutralize cholera toxin and STa toxin and inhibit adherence of K88ac fimbrial E. coli. Clin Vaccine Immunol 17: 1859-1867.
[52] Benz I, Schmidt MA (1992) AIDA-I, the adhesin involved in diffuse adherence of the diarrhoeagenic Escherichia coli strain 2787 (O126:H27), is synthesized via a precursor molecule. Mol Microbiol 6: 1539-1546.
[53] Keller R, Ordonez JG, de Oliveira RR, Trabulsi LR, Baldwin TJ (2002) Afa, a diffuse adherence fibrillar adhesin associated with enteropathogenic Escherichia coli. Infect Immun 70: 2681-2689.
[54] Jiang X, Chai TJ (1996) Survival of Vibrio parahaemolyticus at low temperatures under starvation conditions and subsequent resuscitation of viable, nonculturable cells. Appl Environ Microbiol 62: 1300-1305.
[55] Roszak DB, Colwell RR (1987) Survival strategies of bacteria in the natural environment. Microbiol Rev 51: 365-379.
[56] Abshire KZ, Neidhardt FC (1993) Analysis of proteins synthesized by Salmonella typhimurium during growth within a host macrophage. J Bacteriol 175: 3734-3743.
Abstract
Materials and methods: EAEC (T8) was grown in vitro by mimicking the in vivo pH conditions found in human stomach and intestine.
Results: The lowest pH where EAEC (T8) was able to grow was pH 4.0 in Luria Bertani (LB) media, albeit with a lower growth rate and the bacteria reached log phase in approximately 7h. However, there was no signifi cant diff erence in the growth pattern at mild acidic pH 5.5. In addition, EAEC (T8) grown at pH 4.0, demonstrated an absence of biofi lm formation, clump or pellicle formation and umbrella shaped hemagglutination pattern and within 3h rod-shaped bacteria appeared as coccoid or spheroid forms with average dimensions approximately half the size of bacteria grown at control pH conditions.
Conclusions: Taken together, our data suggest that the changes in the associated phenotypic characters might relate to the adaptation and survival of EAEC under acid stress conditions.
Abbreviations
AA: Aggregative Adherence; AAF: Aggregative Adherence Fimbriae; AR: Acid Resistance; DMEM: Dulbecco’s Modifi ed Eagle’s Medium; EAEC: Enteroaggregative Escherichia coli ; EAEC (T8): Clinical Isolate of EAEC; EDTA: Ethylene Diamine Tetraacetic Acid; EHEC: Enterohemorrhagic Escherichia coli ; HA: Hemagglutination; FCS: Fetal Calf Serum; HEp-2: Human Epithelial Pharyngeal Cell Line; LA: Luria Agar; LB: Luria Bertani; M9: Minimal media; MRHA: Mannose Resistant Hemagglutination; OMP: Outer Membrane Protein; PBS: Phosphate Buff ered Saline; PI: Propidium Iodide